From Duke University, 21/03/23
When a couple of plucky graduate students took the first picture with their pieced-together microscope, it turned out better than they’d hoped.
Sure, there was a hole in one section and another was upside down — but they could still find Waldo.
By the following day, the duo sorted out their software issues and demonstrated a successful proof-of-principle device on the classic children’s puzzle book.
By combining 24 smartphone cameras into a single platform and stitching their images together, they created a single camera capable of taking gigapixel images over an area about the size of a piece of paper.
Six years, several design iterations and one startup company later, the researchers made an unexpected discovery.
Perfecting the process of stitching together dozens of individual cameras with subpixel resolution simultaneously allowed them to see the height of objects too.
“It’s like human vision,” said Roarke Horstmeyer, assistant professor of biomedical engineering at Duke University.
“If you merge multiple viewpoints together (as your two eyes do), you see objects from different angles, which gives you height.”
“When our colleagues studying zebrafish used it for the first time, they were blown away. It immediately revealed new behaviors involving pitch and depth that they’d never seen before.”
In a paper published online March 20 in Nature Photonics, Horstmeyer and his colleagues show off the capabilities of their new high-speed, 3D, gigapixel microscope called a Multi Camera Array Microscope (MCAM).

Whether recording 3D movies of the behavior of dozens of freely swimming zebrafish or the grooming activity of fruit flies at near cellular-level detail across a very wide field of view, the device is opening new possibilities to researchers the world over.
The latest version of MCAM relies on 54 lenses with higher speed and resolution than the prototype that found Waldo. Building upon recent work completed in close collaboration with Dr. Eva Naumann’s lab at Duke, innovative software gives the microscope the ability to take 3D measurements, provide more detail at smaller scales and make smoother movies.
The highly parallelized design of the MCAM, however, creates its own data processing challenges, as a few minutes’ worth of recording can produce over a terabyte of data.
“We’ve developed new algorithms that can efficiently handle these extremely large video datasets,” said Kevin C. Zhou, a postdoctoral researcher in Horstmeyer’s lab and lead author of the paper.”
“Our algorithms marry physics with machine learning to fuse the video streams from all the cameras and recover 3D behavioral information across space and time.”
“We’ve made our code open source on Github for everyone to try out.”
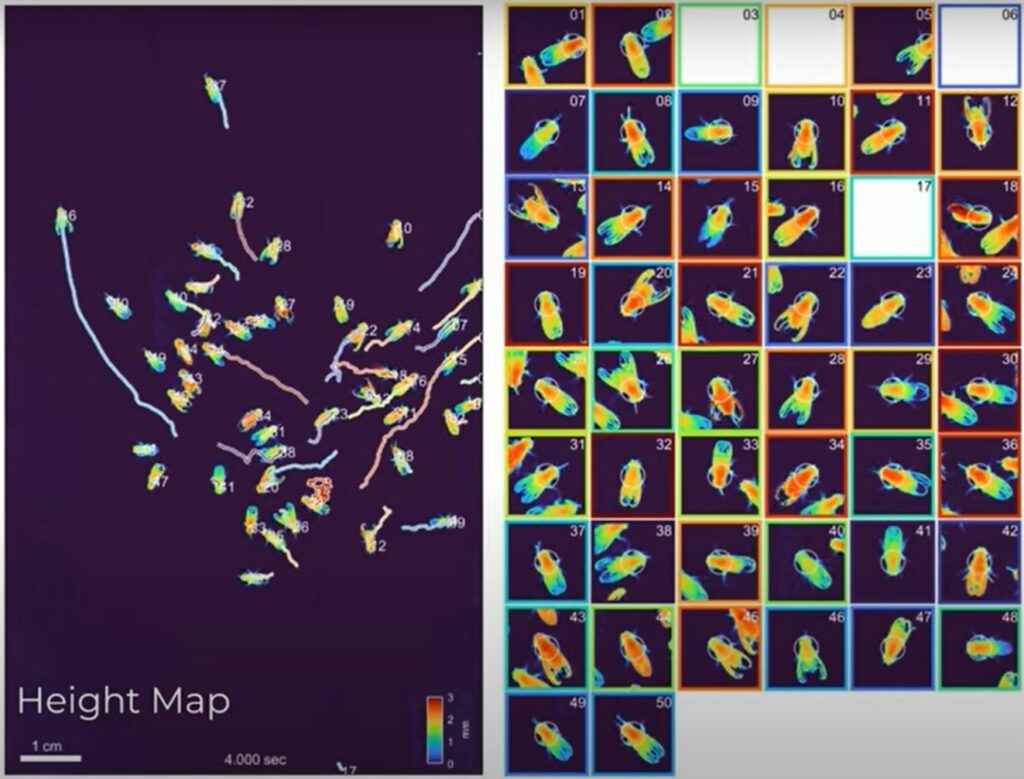
At the University of California – San Francisco, Matthew McCarroll watches the behavior of zebrafish exposed to neuroactive drugs.
By looking for changes in behavior due to different classes of drugs, researchers can discover new potential treatments or better understand existing ones.
In the paper, McCarroll and his group describe interesting movements they’d never seen before thanks to using this camera.
The 3D capabilities of the MCAM, coupled with its all-encompassing view, allowed them to record differences in the fish’s pitch, whether they trended toward the top or bottom of their tanks and how they tracked prey.
“We’ve long been building our own rigs with single lenses and cameras, which have worked well for our purposes, but this is on a whole other level,” said McCarroll, an independent scientist studying pharmaceutical chemistry in the UC system’s professional researcher series.
“We’re just biologists tinkering with optics. It’s incredible to see what a legit physicist can come up with to make our experiments better.”
At Duke, the laboratory of Michel Bagnat, professor of cell biology, also works with zebrafish.
But rather than watching for drug-induced behavioral changes, the researchers study how the animals develop from an egg into a fully formed adult on a cellular level.
In previous studies, the researchers needed to anesthetize and mount the developing fish to keep them steady while measurements were taken with lasers.
But knocking them out for prolonged periods of time might also cause changes in their development that could skew the experiment’s results.
With the help of the new MCAM, the researchers have shown that they’re able to get all of these measurements while the fish live their lives unencumbered, no knockouts or clamps required.
“With the 3D and fluorescent imaging capabilities of this microscope, it could change the course of how a lot of developmental biologists do their experiments,” said Jennifer Bagwell, a research scientist and lab manager in the Bagnat lab.
“Especially if it turns out that anesthetizing the fish affects their development, which is something we’re studying right now.”
Besides tracking entire communities of small animals such as zebrafish in experiments, Horstmeyer hopes this work will also allow for larger automated parallel studies.
For example, the microscope can watch a plate with 384 wells loaded with a variety of organoids to test potential pharmaceutical reactions, recording the cellular responses of each tiny experiment and autonomously flagging any results of interest.
“The modern laboratory is becoming more automated every day, with large well plates now being filled and maintained without ever touching a human hand,” Horstmeyer said.
“The sheer volume of data this is creating demands for new technologies that can help automate the tracking and capturing of the results.”
Along with coauthor Mark Harfouche, who was the brains behind capturing their first image of Waldo, Horstmeyer has launched a startup company called Ramona Optics to commercialize the technology.
One of its early licensers, MIRA Imaging, is using the technology to “fingerprint” fine art, collectables and luxury goods to inoculate against forgery and fraud.
Further examples of the microscope in action can be found at:
- MCAM project webpage: https://mcam.deepimaging.io/
- Sample videos: https://gigazoom.rc.duke.edu/
- Open source code: https://github.com/kevinczhou/3D-RAPID
- Sample raw video files: https://doi.org/10.7924/r4db86b1q
This research is supported by the Office of Research Infrastructure Programs (ORIP), Office of the Director, National Institutes of Health of the National Institutes of Health and the National Institute of Environmental Health Sciences (NIEHS) of the National Institutes of Health (R44OD024879), the National Cancer Institute (NCI) of the National Institutes of Health (R44CA250877), the National Institute of Biomedical Imaging and Bioengineering (NIBIB) of the National Institutes of Health (R43EB030979), the National Science Foundation (2036439) and a Duke Coulter Translational Partnership Award.
Links
https://www.nature.com/articles/s41566-023-01171-7



